嗯,用户让我帮忙总结这篇文章的内容,控制在100字以内,而且不需要特定的开头。首先,我需要通读整篇文章,抓住主要观点。 文章主要讨论了自然语言处理中的实体识别(NER)问题,特别是标注数据的成本高昂。传统的做法需要大量的人工标注,尤其是对于专业领域如医疗和金融,这不仅耗时还昂贵。作者介绍了一种新的方法,利用大语言模型(LLM)进行标注,从而减少对人工标注的依赖。 接下来,文章通过两个案例研究展示了这种方法的有效性。一个案例是NCBI疾病语料库,另一个是临床EHR中的健康ID。结果显示,在结构清晰的实体类型上,LLM标注可以达到生产级别的性能,而无需大量的人工标注。 总结起来,文章的核心是用LLM替代大部分人工标注工作,从而降低NER模型开发的时间和成本。用户可能需要一个简洁明了的摘要来快速了解文章内容。 </think> 文章讨论了自然语言处理中实体识别(NER)的人工标注成本高昂问题,并提出了一种利用大语言模型(LLM)进行自动标注的新方法。通过案例研究(如NCBI疾病语料库和临床EHR数据),展示了LLM在减少人工标注需求的同时仍能实现高质量模型的效果。该方法将繁琐的标注过程简化为少量验证集和规则编写,大幅降低了时间和经济成本。 2026-3-24 19:12:20 Author: securityboulevard.com(查看原文) 阅读量:7 收藏
The problem: human annotation is expensive
Ever since BERT, NLP practitioners have enjoyed a golden age of text classification — simply download your favorite pre-trained model from HuggingFace and fine-tune on training data for any number of text classification problems. This has allowed practitioners to set aside fun games of eeking out architectural improvements and focus on the highest leverage move: collecting high quality training and evaluation data. Unfortunately, this has been a fundamentally slow and expensive process. Aligning human annotators to guidelines, ensuring high quality work, and just sitting down and looking at the data are all slow, difficult steps.
As an example of the cost and effort to develop a high quality training corpus, consider the NCBI Disease Corpus, a standard benchmark for biomedical NER. Its creators employed 14 annotators with biomedical informatics backgrounds across two summers (2011 and 2012), each labeling ~120 PubMed abstracts. Every document was independently annotated by two people, disagreements were discussed to reach consensus, and then a final pass checked consistency across the entire corpus. The result, 793 annotated abstracts, required hundreds of person-hours of expert work.
This is the norm in NER. Building a high-quality training corpus means recruiting domain experts, developing annotation guidelines through iterative pilot rounds, running multi-annotator campaigns, adjudicating disagreements, and auditing for consistency. For specialized domains like healthcare or finance, qualified annotators are scarce and expensive. The annotation bottleneck is often the single biggest barrier to deploying custom NER.
What if you could skip most of that?
Our approach: LLM annotations + lightweight fine-tuning
Tonic Textual‘s custom entity workflow replaces the human annotation campaign with an LLM. The process:
- Write annotation guidelines in natural language: what the entity is, what to include, what to exclude.
- Upload a small validation set with ground truth labels (as few as 10-100 documents).
- Iterate on guidelines: the system compares LLM annotations to your ground truth and suggests improvements.
- Upload unlabeled training data. The LLM annotates it using your refined guidelines.
- Train a lightweight encoder model on the LLM annotations for fast, cheap inference at deployment.
The expensive human annotation is confined to a small validation set. The bulk training data is annotated by the LLM, which follows your guidelines consistently across thousands of documents.
Case study: NCBI disease corpus
We tested this on the NCBI Disease Corpus: 593 training documents, 100 validation, 100 held-out test. All PubMed abstracts annotated with disease mentions, including genuinely hard cases like abbreviations that refer to both genes and diseases (“APC”, “VHL”), disease names used as modifiers (“HD gene”), and composite mentions (“colorectal and ovarian cancers”).
The annotation challenge
In biomedical text, abbreviations like “APC,” “VHL,” and “DM” name both a gene and its associated disease. The NCBI corpus labels these as disease mentions even when they appear in gene context: “the APC gene,” “DM protein kinase,” “A-T allele” all contain disease spans. An NER model has to learn to tag “APC” as a disease in “the APC gene caused APC” while a naive model would suppress tagging near words like “gene,” “kinase,” and “allele.” Our error analysis found this gene/disease polysemy accounts for 60% of complete misses.
How much do human labels actually help?
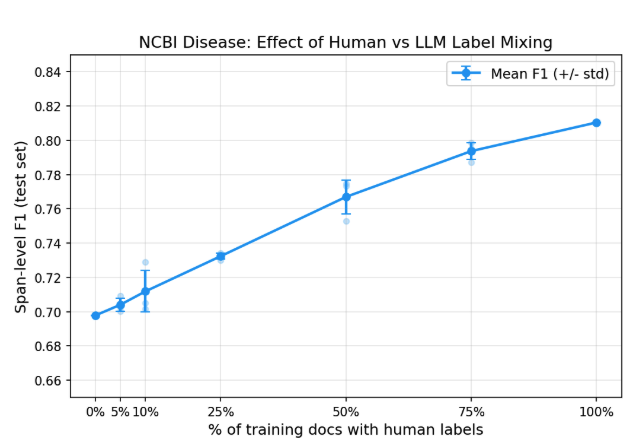
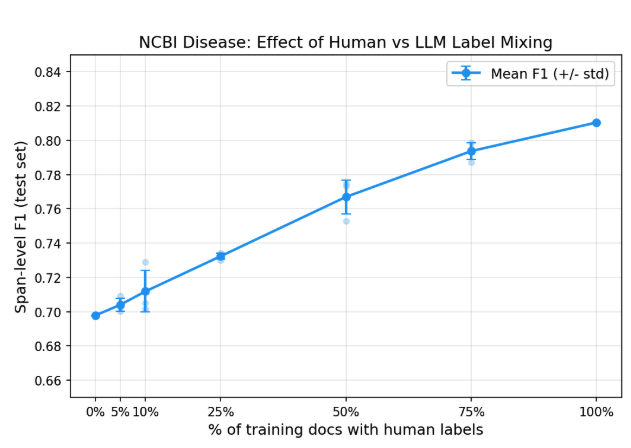
To answer this directly, we ran a controlled experiment. We took the LLM annotations produced by the custom entity workflow (F1=0.71 against ground truth on the training set) and trained RoBERTa LoRA models in our own training loop so we could precisely control the label mix. All 593 training documents were always used; we varied what fraction had their LLM labels replaced with human ground truth.
| % Human Labels | # Human-Labeled Docs | Test F1 |
|---|---|---|
| 0% | 0 | 0.70 |
| 5% | 30 | 0.70 |
| 10% | 59 | 0.71 |
| 25% | 148 | 0.73 |
| 50% | 297 | 0.77 |
| 75% | 445 | 0.79 |
| 100% | 593 | 0.81 |
With zero human-labeled training data, LLM annotations alone produce a model at 0.70 F1. Adding human labels helps (the full human-labeled set reaches 0.81) but the returns diminish steeply. The first 148 human-labeled documents (25%) adds only 3 F1 points. Each subsequent increment adds less.
The cost of NER is no longer “annotate all your training data.” It’s “annotate a good validation set, write good guidelines, and let the LLM handle the rest.”
Putting the numbers in context
How does 0.70 F1 compare? We benchmarked several approaches on the same test set:
| Approach | Test F1 | Human Annotation Required |
|---|---|---|
| GLiNER2 zero-shot | 0.395 | None |
| Custom entity workflow (Textual SDK) | 0.71 | Validation set only (~10 docs) |
| Same training loop, 100% human labels | 0.81 | Full training set (593 docs) |
GLiNER2, a recent zero-shot NER model, achieves high precision (0.82) but misses three-quarters of disease mentions (recall = 0.26). It catches obvious cases like “colorectal cancer” but misses abbreviations, modifiers, and disease classes. Zero-shot NER on specialized domains remains unreliable.
Meanwhile, the Textual SDK workflow using just 10 human-labeled documents for guideline refinement achieves 0.71 F1, matching the 0% human learning curve result. The human effort that matters isn’t labeling thousands of training examples; it’s writing precise guidelines and validating them against a curated set.
The complexity factor
We should be honest: 0.70-0.81 F1 is below the ~0.87-0.90 that specialized biomedical models (BioBERT, PubMedBERT) achieve with full fine-tuning on this benchmark. Two factors explain most of the gap:
- We used
roberta-base, a general-purpose model. A biomedical base model would close much of this gap. - Gene/disease polysemy is uniquely hard in this corpus. Abbreviations like APC and DM get labeled as diseases even in genetic context (“the APC gene”). Our error analysis found this accounts for 60% of complete misses.
Both are solvable (swap the base model, refine the guidelines) but beside the point. This benchmark isn’t about setting a new SOTA. It’s about showing that the annotation bottleneck no longer blocks training useful NER models.
What this means in practice?
The traditional NER pipeline looks like:
Recruit annotators -> Develop guidelines -> Pilot annotation -> Adjudicate -> Full annotation campaign -> Train model -> Evaluate -> Iterate
This takes weeks or months of calendar time and thousands of dollars in annotation costs per entity type.
The custom entity workflow compresses this to:
Write guidelines -> Label ~100 validation examples -> Refine guidelines -> Train -> Evaluate
The total human effort is a validation set of 10-100 documents, plus the intellectual work of writing good guidelines. The LLM annotates hundreds or thousands of training documents in minutes instead of weeks.
Our two case studies bracket the range of what to expect. On a difficult academic benchmark with context-dependent labels and gene/disease ambiguity, LLM annotations alone produce a 0.70 F1 model, and mixing in human labels pushes steadily toward 0.81. On a practical production entity type with clear structural patterns, the workflow exceeds production release thresholds on the first try.
The cost of performance is now concentrated where it should be: understanding what you want to extract and validating that understanding against a high-quality dataset.
Case study: EHR healthcare IDs
The NCBI benchmark tests a hard public dataset, but there’s an elephant in the room. The NCBI Disease Corpus is well-known in NLP, and the LLM annotator has almost certainly seen it (or data very like it) during pretraining. How much of the 0.70 F1 comes from the workflow, and how much from the LLM recognizing familiar examples?
To control for this, we also evaluated on a private dataset that cannot have leaked into any LLM’s training data: healthcare identifiers (MRNs, account numbers, unit/bed numbers, encounter IDs) in clinical EHR documents.
- Validation set: 123 documents, 266 healthcare_number spans (human-labeled)
- Training set: 1,119 documents (unlabeled, LLM annotates)
Guideline refinement
Starting from our existing production guidelines for healthcare_number, the LLM refinement loop rapidly improved annotation quality:
| Iteration | LLM F1 |
|---|---|
| v1 (seed) | 0.896 |
| v2 | 0.946 |
| v3 | 0.960 |
Three iterations, each taking a few minutes. The system identified edge cases (distinguishing healthcare IDs from procedure codes like CPT and ICD, handling alphanumeric formats like “482-A”) and tightened the guidelines automatically.
Trained model
The encoder model trained on 1,119 LLM-annotated documents achieved 0.947 F1, exceeding our production release threshold of 0.914.
| Metric | Custom Entity Model | Production Threshold |
|---|---|---|
| F1 | 0.947 | 0.914 |
| Precision | – | 0.900 |
| Recall | – | 0.929 |
This is a production-ready model, trained with zero human-labeled training data. The only human effort was the 123-document validation set, which we already had from our existing annotation pipeline.
What this tells us
The NCBI benchmark is intentionally hard: gene/disease polysemy, subtle context-dependent labels, a corpus designed to stress-test NER systems. Healthcare IDs are a more typical production entity type. They are structurally distinctive (numbers, alphanumeric codes), appear in predictable contexts (after “MRN:”, “Account#:”), and have clear inclusion/exclusion criteria.
For entity types like this, where the surface patterns are distinctive and the guidelines are relatively unambiguous, the workflow delivers production-quality models with zero human-labeled training data.
Ready to train your own custom NER model without the annotation bottleneck? Tonic Textual’s custom entity workflow lets you go from guidelines to a production-ready model in hours, not months. Start your free trial and see how far you can get with just a small validation set.
如有侵权请联系:admin#unsafe.sh